New study highlights major step forward in monitoring ocean health
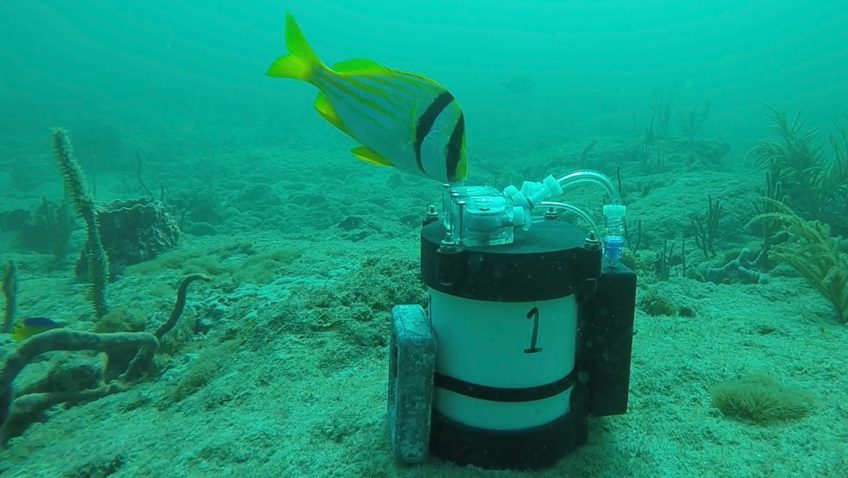
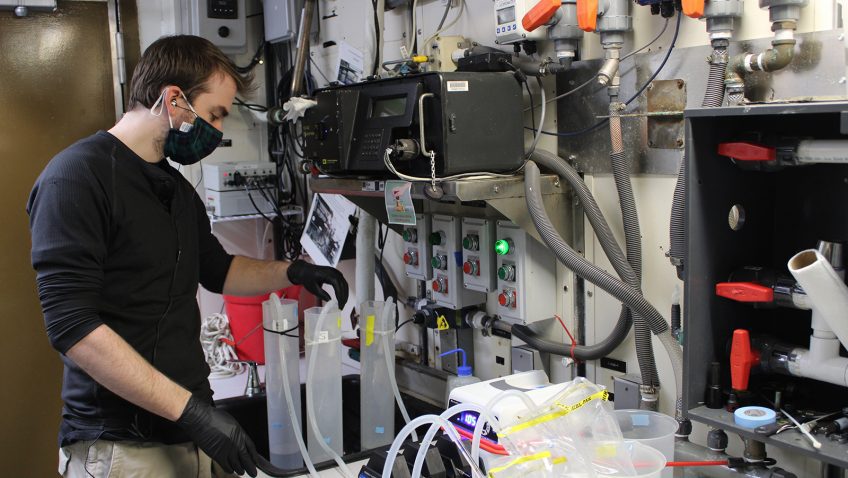
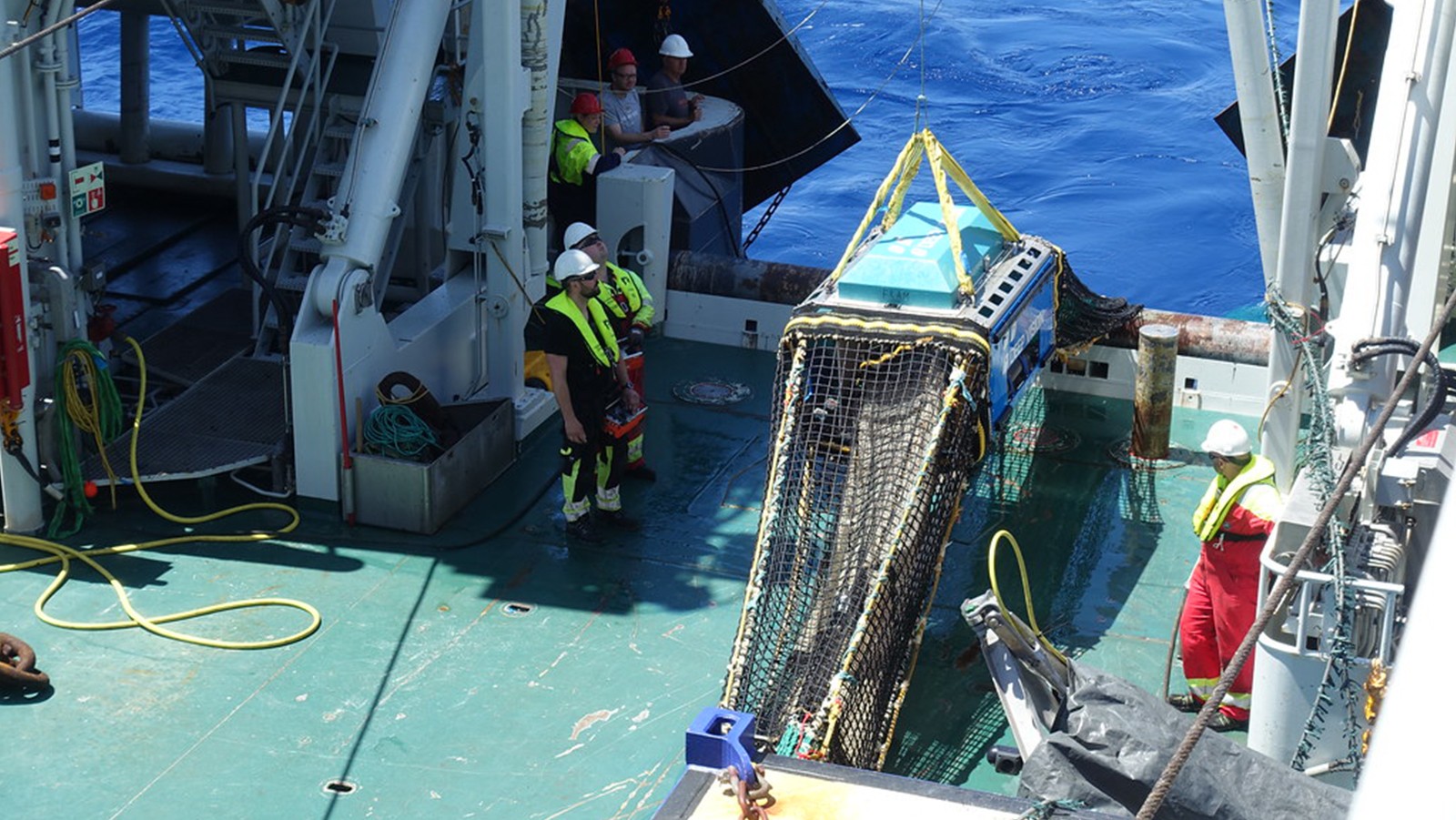
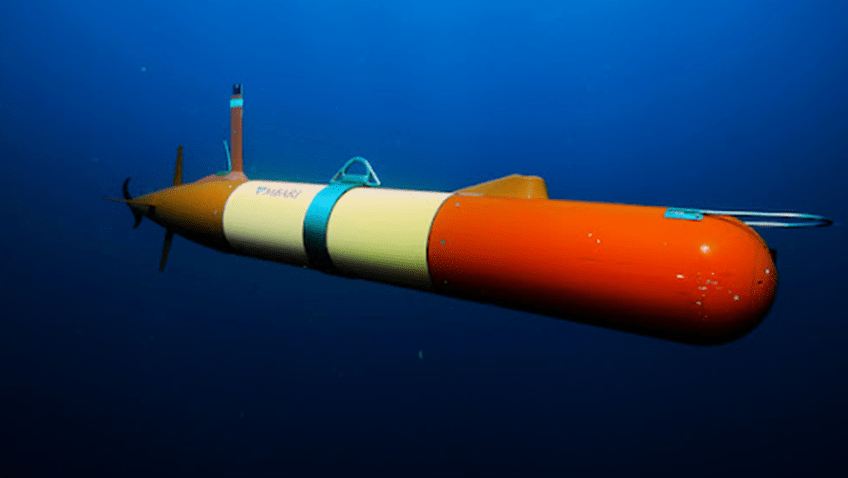
In a major step forward for monitoring the biodiversity of marine systems, a new study published in Environmental DNA details how Monterey Bay Aquarium Research Institute (MBARI) and NOAA’s Atlantic Oceanographic & Meteorological Laboratory (AOML) researchers are using autonomous underwater robots to sample environmental DNA (eDNA). eDNA allows scientists to detect the presence of aquatic species from the tiny bits of genetic material they leave behind. This DNA soup offers clues about biodiversity changes in sensitive areas, the presence of rare or endangered species, and the spread of invasive species—all critical to understanding, promoting, and maintaining a healthy ocean.